![Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ] Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7755/1/fig-1-2x.jpg)
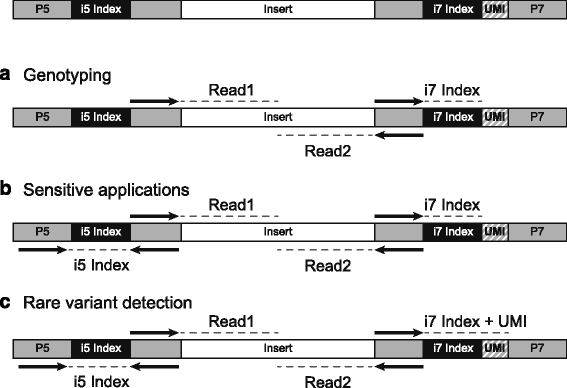
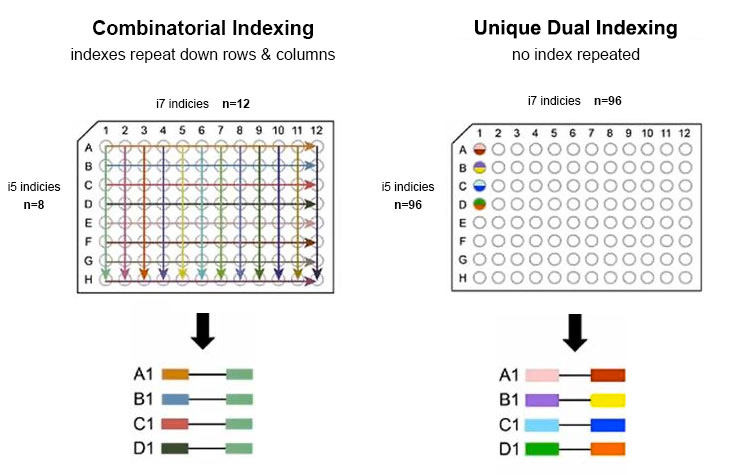
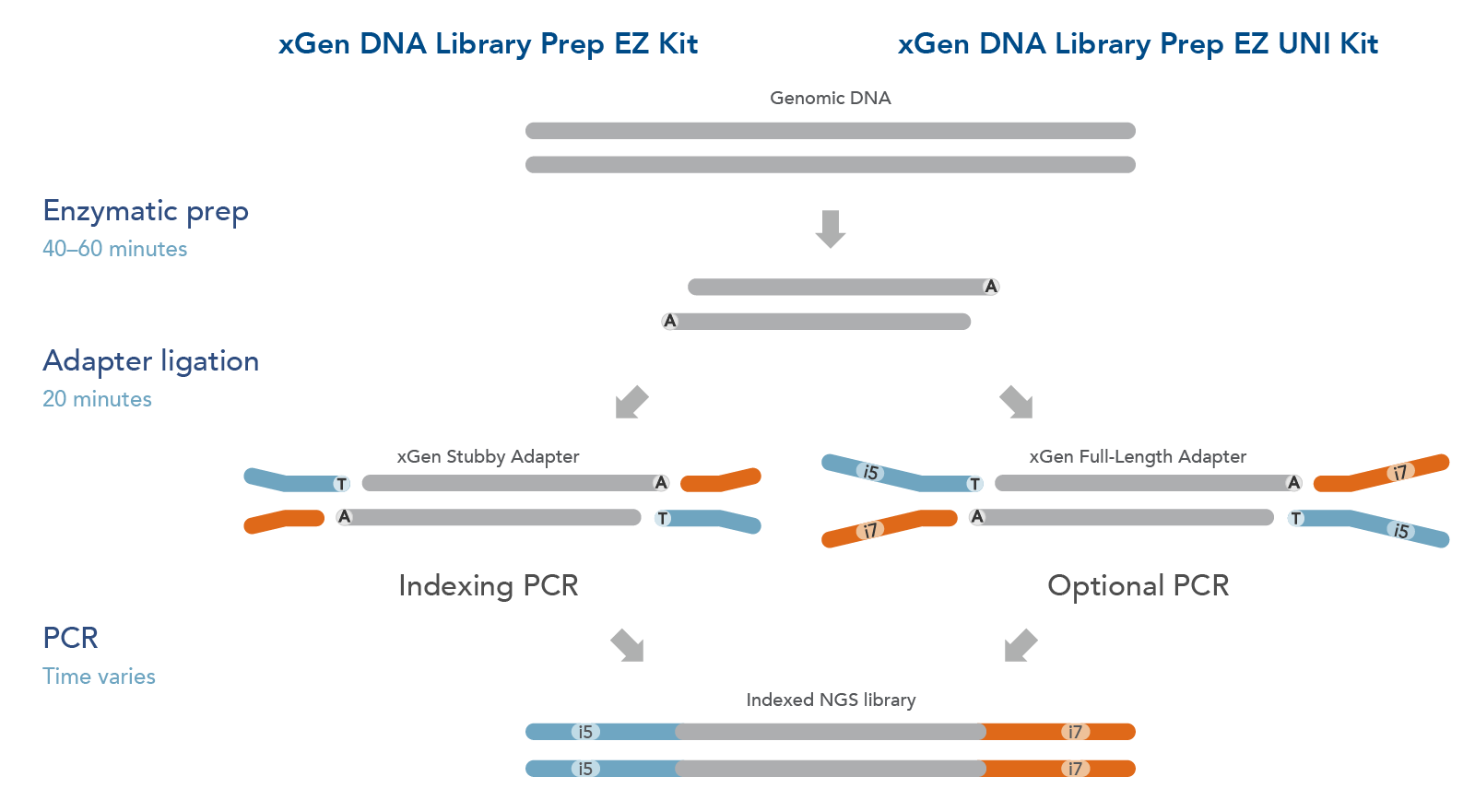
Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]
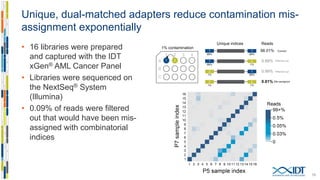
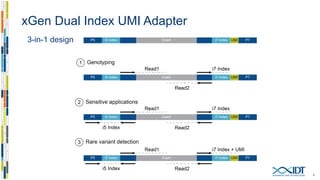
Dual index adapters with UMIs resolve index hopping and increase sensitivity of variant detection on Vimeo
![Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ] Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2019/7755/1/fig-4-2x.jpg)
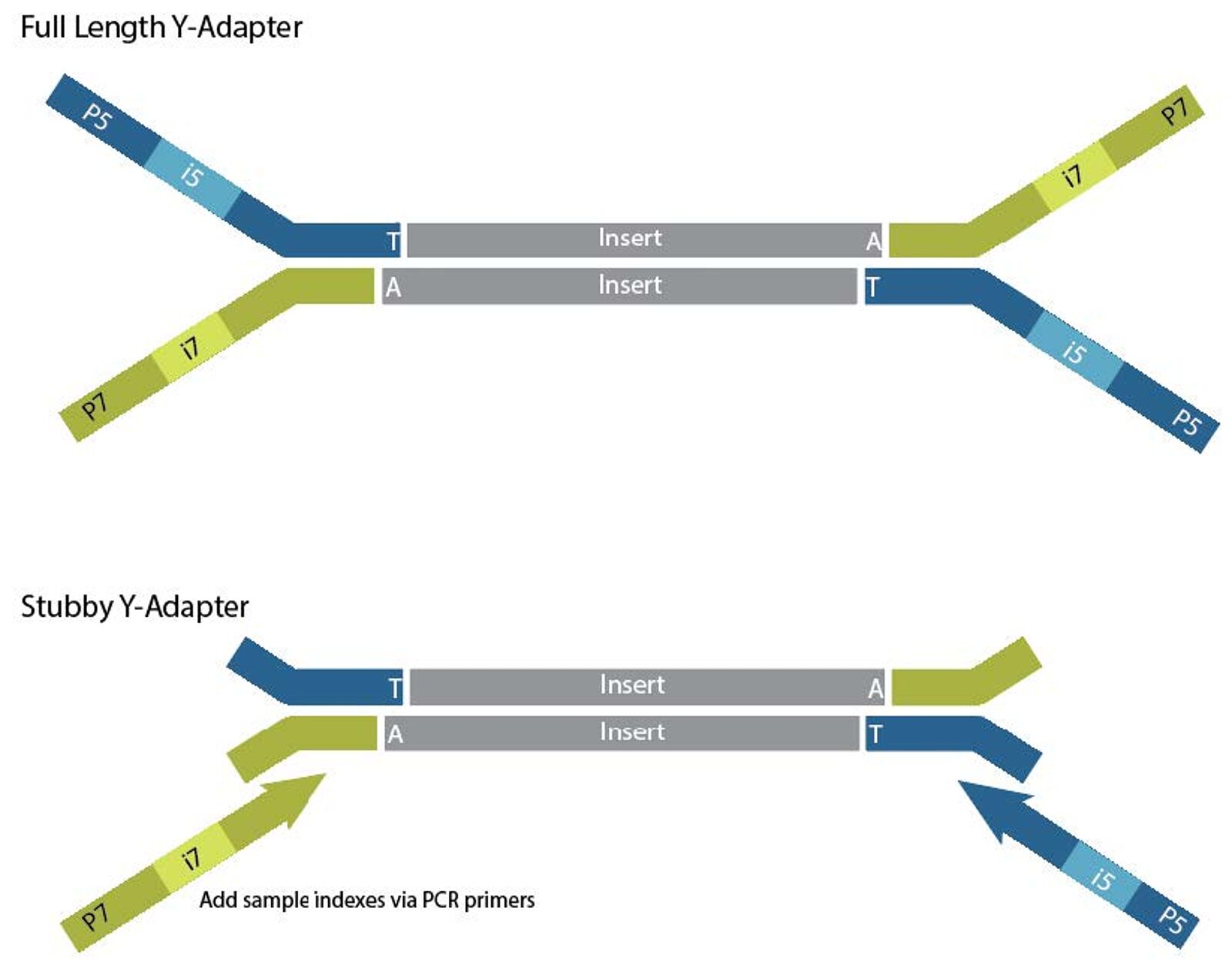
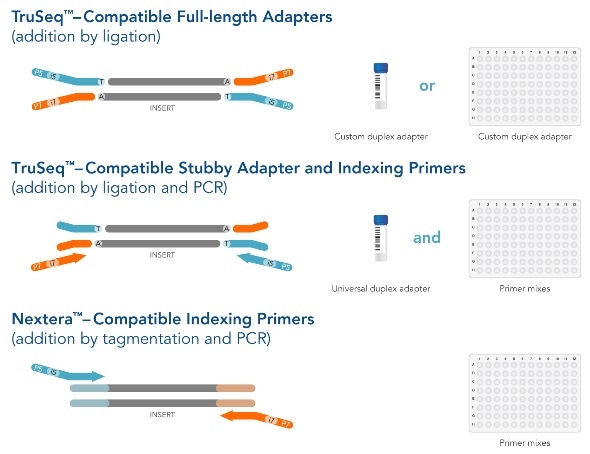
Adapterama I: universal stubs and primers for 384 unique dual-indexed or 147,456 combinatorially-indexed Illumina libraries (iTru & iNext) [PeerJ]
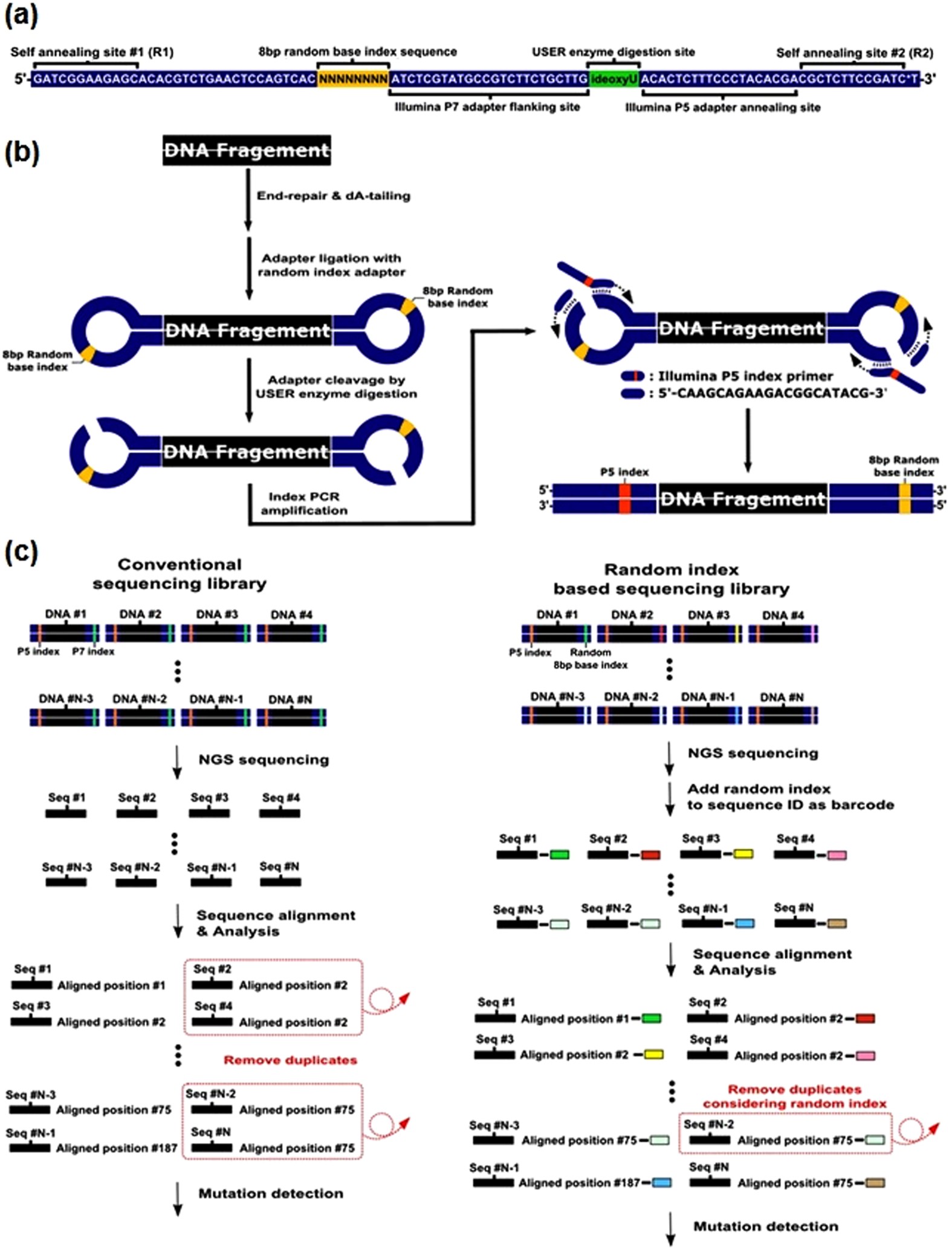
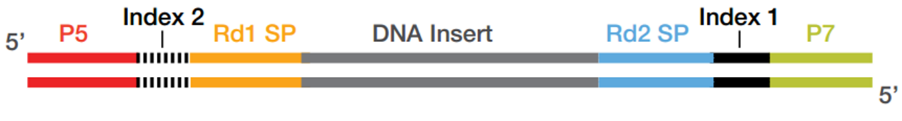
Asymmetrical barcode adapter-assisted recovery of duplicate reads and error correction strategy to detect rare mutations in circulating tumor DNA | Scientific Reports